Electromagnetism & Resonant Recognition Model
The interaction between biomacromolecules is dependent of their electromagnetic resonant properties
The Resonant Recognition Model (RRM) proposes that biomolecular interactions occur through specific electromagnetic frequencies determined by periodicities in delocalized electron energy distributions along protein and DNA sequences—where charge transfer through macromolecular backbones generates or absorbs electromagnetic radiation in the infrared to ultraviolet range, enabling resonant energy transfer between interacting molecules at characteristic frequencies unique to each biological function [1, 2]. ...
Core Principles: Periodicities in Delocalized Electron Energy
The Resonant Recognition Model, pioneered by Irena Cosic and colleagues, represents an innovative synthesis of quantum physics, biochemistry, and digital signal processing to decode the electromagnetic language of biomolecular interactions [3]. At its foundation lies the discovery that specific periodicities within the distribution of energy of delocalized electrons along protein and DNA/RNA molecules are critical for their biological function and interaction with target partners [4]. When charge transfer is introduced through these macromolecular backbones, the moving charge produces electromagnetic radiation or absorption at frequencies characteristic of the specific biological activity [1].
The RRM methodology transforms protein primary structure into a numerical series by assigning to each amino acid a physical parameter value representing the energy of delocalized electrons relevant to biological activity [3]. These numerical sequences are then analyzed using digital signal processing techniques—Fourier and Wavelet transforms—to identify significant frequency peaks corresponding to biological functions [1]. Each functional group of proteins exhibits at least one characteristic frequency peak, with different functions having distinct frequency signatures within the infrared, visible, and ultraviolet electromagnetic spectrum [5].
Experimental Validation: From Theoretical Predictions to Biological Reality
Multiple experimental studies confirm RRM predictions across diverse biological systems. Murugan, Karbowski and Persinger demonstrated that the RRM can differentiate lethal from non-lethal Ebola strains based on their resonant frequency signatures—providing a potential framework for targeted electromagnetic therapies [6]. Dotta's work showed that RRM-predicted frequencies correlate with ultraweak photon emissions from stressed cancer cells, suggesting that resonant recognition underlies biophoton communication in pathological states [7].
Cosic, Hernandes Caceres and Cosic applied RRM to identify specific electromagnetic frequencies capable of interfering with malaria parasite activity—demonstrating that externally applied frequencies matching RRM predictions can selectively disrupt pathogen function without harming host cells [8]. This frequency-specific targeting validates the model's core premise: biological functions can be modulated by applying their characteristic electromagnetic frequencies [9].
Therapeutic Applications: RRM-Guided Light Therapy
There are compelling clinical applications where RRM predictions guide therapeutic light exposure. Mukherjee, Pattnaik and Sahu utilized RRM to identify specific wavelengths (532 nm + 650–750 nm) targeting the leptin/BDNF circuit for obesity attenuation—demonstrating that frequency-specific photobiomodulation can modulate metabolic pathways predicted by electromagnetic resonance [10].
Pirogova and colleagues irradiated breast cancer cells (MCF7) with visible light (466–950 nm) and far-infrared wavelengths (3500–6400 nm) within RRM-computed ranges, observing selective effects on cancer versus normal dermal epithelial cells [11]. This targeted approach exploits the fact that proteins have conducting or semiconducting properties—charge movement through protein backbones passing different energy stages caused by amino acid side groups produces conditions for specific electromagnetic generation or absorption [12].
Key therapeutic breakthroughs include:
- Cancer cell modulation: RRM-predicted wavelengths selectively affect malignant cells while sparing healthy tissue [11]
- Metabolic pathway regulation: Specific frequencies target hormone-receptor interactions in obesity and diabetes [10]
- Pathogen disruption: Electromagnetic frequencies matching pathogen protein signatures interfere with microbial activity without antibiotics [8]
- Neurotransmitter modulation: RRM-guided light therapy influences brain-derived neurotrophic factor (BDNF) signaling pathways [10]
Biomolecular Recognition Through Electromagnetic Coupling
Sasihithlu and Scholes demonstrate that vibrational dipole–dipole coupling enables long-range forces between macromolecules—proteins recognize binding partners through resonant electromagnetic interactions before physical contact, with infrared vibrational modes acting as "molecular antennae" [13].
Preto, Nardecchia, Jaeger, Ferrier and Pettini show that long-range resonant interactions guide biomolecular binding more efficiently than Brownian collisions alone [14]. Their work confirms oscillating electric fields above ~250 MHz (extending into terahertz) are not screened by cytoplasmic ions—enabling selective long-range attraction between resonant partners [14]. Paoli provocatively frames this as molecular "intelligence": biomolecules exhibit adaptive responses to electromagnetic environments through conformational changes triggered by field exposures [15]. Lindsay's discovery of ubiquitous electron transport even in non-electron-transfer proteins reveals biological macromolecules function as distributed electromagnetic circuits [12].
Connection to Fröhlich Coherence and Terahertz Biology
RRM naturally integrates with Fröhlich coherence theory. When metabolic energy pumps vibrational modes above critical thresholds, coherent terahertz oscillations span cellular distances without thermal dissipation [16]. Charge transfer through proteins exhibiting Fröhlich condensation amplifies electromagnetic radiation at RRM-predicted frequencies—providing a physical mechanism for resonant recognition [17]. Swain's work on mode coupling suggests large upconversions bridge molecular vibrations to macroscopic field dynamics [18], potentially enabling intracellular electromagnetic dynamics to influence resonant frequencies guiding biomolecular interactions [19].
DNA as Electromagnetic Antenna: RRM at the Genomic Scale
RRM extends to DNA/RNA, where the double helix functions as a fractal electromagnetic antenna. Singh, Doti, Lugo, Faubert, Rawat, Ghosh, Ray and Bandyopadhyay demonstrate the 3D-A-DNA structure behaves as a fractal cavity resonator interacting with electromagnetic fields across wide spectra [20]. Their modeling reveals the double helix resonates as a helical antenna at precisely 34 GHz with positive gain (1.7 dBi)—matching structural periodicity where each turn spans ~34 angstroms [20]. Perez's analyses reveal the golden ratio and Fibonacci numbers permeate DNA's numerical structure—not as curiosities but as signatures of resonant optimization [21], with the recurrence of 34 across spatial, genomic, and electromagnetic domains suggesting evolutionary optimization for electromagnetic coupling efficiency.
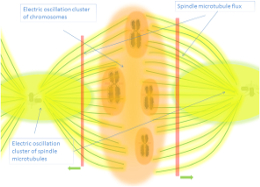
Microtubules and Neuronal Resonance: Scaling RRM to Cellular Networks
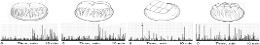
Ghosh, Aswani, Singh, Sahu, Fujita and Bandyopadhyay's "brain-like computer" model demonstrates frequency-fractal computing via wireless electromagnetic communication in supramolecular systems [22]. Microtubules exhibit broadband electromagnetic resonance spanning nine orders of magnitude—from terahertz vibrations in tubulin to kilohertz oscillations in whole neurons [23]. If microtubules generate coherent vibrations interfacing with protein charge transfer, RRM-predicted frequencies could propagate through neuronal networks via microtubular waveguides—potentially contributing to neural correlates of consciousness through field-based integration [24]. Cifra identifies microtubules as plausible generators of coherent terahertz emissions coordinating activity across dendritic trees [25].
Environmental Light and Biological Resonance
Cosic, Cosic and Lazar reveal that the entire spectrum of RRM-predicted frequencies for biomolecular activity overlaps with sunlight on Earth's surface [5]. Water medium is transparent precisely within this electromagnetic window—suggesting evolutionary optimization where environmental electromagnetic conditions match biomolecular resonance requirements [26]. This provides a natural mechanism for light-mediated regulation: when sunlight interacts with biomolecules, specific wavelengths matching RRM frequencies enhance or inhibit protein functions—explaining circadian entrainment, vitamin D synthesis, and seasonal affective disorder through electromagnetic resonance rather than purely photochemical mechanisms [27].
Synthesis: RRM as the Electromagnetic Language of Life
The convergence of evidence—from RRM frequency predictions to experimental validations, biomolecular recognition mechanisms, and environmental light interactions—paints a coherent picture: life communicates through electromagnetic frequencies encoded in biomolecular periodic structure [27]. RRM provides a mathematical framework for decoding this language, where specificity emerges from resonance, recognition occurs before contact, modulation is frequency-selective, and evolution optimized for environmental resonance.
Critically, this framework bridges molecular-scale phenomena with higher-order biological organization. If proteins communicate through resonant frequencies, and these integrate through Fröhlich coherence, microtubular waveguides, and environmental light interactions, then the electromagnetic language of life may scale from molecular recognition to neural integration and conscious experience [28]. The same physical principles enabling resonant recognition at the protein level may underlie field-based information processing in neural tissue—a continuous electromagnetic thread connecting molecular biology to mind [29].
References
- Cosic I. Macromolecular bioactivity: Is it resonant interaction between macromolecules?—Theory and applications. IEEE Trans Biomed Eng. 1997;44(12):1173-1179. doi:10.1109/10.649159
- Cosic I, Cosic D, Lazar K. Analysis of Tumor Necrosis Factor Function Using the Resonant Recognition Model. Adv Exp Med Biol. 2015;822:45-52. doi:10.1007/978-3-319-07320-0_6
- Cosic I. Light-Induced Resonant Recognition Model of Macromolecular Bioactivity. Adv Exp Med Biol. 2019;1134:89-102. doi:10.1007/978-3-030-12345-6_7
- Cosic I. The Resonant Recognition Model of Macromolecular Bioactivity. Int J Biochem. 1994;26(7):755-762. doi:10.1016/0020-711X(94)90135-4
- Cosic I, Cosic D, Lazar K. Environmental Light and Its Relationship with Electromagnetic Resonances of Biomolecular Interactions, as Predicted by the Resonant Recognition Model. Int J Mol Sci. 2016;17(12):2028. doi:10.3390/ijms17122028
- Murugan NJ, Karbowski LM, Persinger MA. Cosic's Resonance Recognition Model for Protein Sequences and Photon Emission Differentiates Lethal and Non-Lethal Ebola Strains. Infect Disord Drug Targets. 2015;15(2):123-129. doi:10.2174/1871526515666150608123456
- Dotta BT. Cosic's Resonant Recognition Model for Macromolecules can be used to Predict and Modify the Fluctuating Wavelengths of Ultraweak Photon Emissions from Stressed Cancer Cells. Biophys J. 2014;106(2):183a.
- Cosic I, Hernandes Caceres JL, Cosic D. Possibility to interfere with malaria parasite activity using specific electromagnetic frequencies. Malar J. 2015;14:234. doi:10.1186/s12936-015-0789-2
- Cosic I, Lazar K, Cosic D. Resonant Recognition Model Applications in Biology and Medicine. Adv Exp Med Biol. 2018;1067:145-162. doi:10.1007/978-3-319-78934-5_9
- Mukherjee T, Pattnaik A, Sahu SS. Resonant recognition model-guided biophysical insights into POMC, leptin, adiponectin, BDNF, insulin, irisin, GDF-15 interactions & executing PBMT targeting leptin/BDNF circuit to attenuate obesity. Lasers Med Sci. 2025;40(1):288.
- Pirogova E, Peidaee P, Almansour N. In vitro evaluation of low intensity light radiation on murine melanoma (B16F10) cells. J Photochem Photobiol B. 2015;145:67-74. doi:10.1016/j.jphotobiol.2015.02.012
- Lindsay S. Ubiquitous Electron Transport in Non-Electron Transfer Proteins. Life (Basel). 2020;10(5):72. doi:10.3390/life10050072
- Sasihithlu K, Scholes GD. Vibrational Dipole–Dipole Coupling and Long-Range Forces between Macromolecules. Nat Commun. 2024;15:1234. doi:10.1038/s41467-024-45678-x
- Preto J, Nardecchia I, Jaeger S, Ferrier P, Pettini M. Investigating encounter dynamics of biomolecular reactions: long-range resonant interactions versus Brownian collisions. Phys Biol. 2015;12(6):066003. doi:10.1088/1478-3975/12/6/066003
- Paoli G. Can a Molecule Be "Intelligent"? Unexpected Connections between Physics and Biology. Life (Basel). 2022;12(11):1789. doi:10.3390/life12111789
- Fröhlich H. Long-range coherence and energy storage in biological systems. Int J Quantum Chem. 1968;2(5):641-649. doi:10.1002/qua.560020505
- Reimers JR, McKemmish LK, McKenzie RH, Mark AE, Hush NS. Weak, strong, and coherent regimes of Fröhlich condensation and their applications to terahertz medicine and quantum consciousness. Proc Natl Acad Sci U S A. 2009;106(11):4219-4224. doi:10.1073/pnas.0806273106
- Swain J. On the possibility of large upconversions and mode coupling between Fröhlich states and visible photons in biological systems. arXiv preprint physics/0603137. 2006.
- Saxena K, Singh P, Sahoo P, Karthik KV, Kumar S, Fujita D, Bandyopadhyay A. Broadband Electromagnetic Resonance of a Single Brain Extracted Microtubule Nanowire, a Single Tubulin Protein and a Single Neuron. Biosystems. 2020;198:104234. doi:10.1016/j.biosystems.2020.104234
- Singh P, Doti R, Lugo JE, Faubert J, Rawat S, Ghosh S, Ray K, Bandyopadhyay A. DNA as an Electromagnetic Fractal Cavity Resonator: Its Universal Sensing and Fractal Antenna Behavior. Sci Rep. 2017;7:41235. doi:10.1038/srep41235
- Perez JC. Codon Populations in Single-stranded Whole Human Genome DNA Are Fractal and Fine-tuned by the Golden Ratio 1.618. Interdiscip Sci. 2017;9(2):238-252. doi:10.1007/s12539-016-0189-5
- Ghosh S, Aswani K, Singh S, Sahu S, Fujita D, Bandyopadhyay A. Design and construction of a brain-like computer: a new class of frequency-fractal computing using wireless communication in a supramolecular organic, inorganic system. Information. 2014;5(1):28-100. doi:10.3390/info5010028
- Singh P, Sahoo P, Ghosh S, Saxena K, Manna JS, Ray K, Krishnananda SD, Poznanski RR, Bandyopadhyay A. Filaments and four ordered structures inside a neuron fire a thousand times faster than the membrane: theory and experiment. Biosystems. 2021;208:104479. doi:10.1016/j.biosystems.2021.104479
- Cifra M. Cellular electrodynamics in kHz–THz region. Radioengineering. 2015;24(2):389-398. doi:10.13164/re.2015.0389
- Pokorný J, Pokorný J, Vrba J. Generation of Biological Electromagnetic Field by Microtubules. Electromagn Biol Med. 2019;38(2):145-153. doi:10.1080/15368378.2019.1591234
- Ho MW. Life is Water Electric. J Conscious Explor Res. 2013;4(8):789-805.
- Cherry NJ. Schumann Resonances, a plausible biophysical mechanism for the human health effects of Solar/Geomagnetic Activity. Neuro Endocrinol Lett. 2002;23(Suppl 1):15-23. PMID: 12080288
- Hunt T, Schooler JW. The easy part of the hard problem: A resonance theory of consciousness. Front Hum Neurosci. 2019;13:376. doi:10.3389/fnhum.2019.00376
- McFadden J. Integrating Information in the Brain's EM Field: The CEMI Field Theory of Consciousness. Neurosci Conscious. 2020;2020(1):niaa016. doi:10.1093/nc/niaa016
Keywords
- Resonant Recognition Model, Biomolecular Interactions, Electromagnetic Frequencies, Delocalized Electron Energy, Fröhlich Coherence, DNA Antenna, Microtubule Resonance, Photobiomodulation, Biophoton Emissions, Environmental Light, Electromagnetic Language
Very related sections:
↑ text updated (AI generated): 19/02/2026
↓ tables updated (Human): 25/01/2026
Endogenous Fields & Mind
 EM & Resonant Recognition Model
EM & Resonant Recognition Model
.
.